![]() By Maciamo Hay,
By Maciamo Hay,
on 16 January 2016 (updated on 29 May 2018)
How to interpret your microbiome results?
Companies like uBiome now make it possible to know what bacteria live in our nose, mouth, gut or skin and help us diagnose potential health issues before they even arise. With genetic testing, microbiome testing is the most revolutionary tool in modern, personalized medicine. But what does it all mean?
Introduction
The human body hosts hundreds of billions of bacteria, a number that varies with the food we eat, how often we wash ourselves, brush our teeth, take antibiotics or drink alcohol or other antibacterials (ginger, garlic, turmeric), but generally exceeds the number of human cells, and can even occasionally outnumber them by a factor of ten to one. These bacteria live in symbiosis with us, helping us digesting foods, metabolizing vitamins, sugars, fats and amino acids, enhancing our immune system, and protecting us from other pathogenic bacteria or fungi. Together with viruses and fungi that reside inside us they form our microbiome. Disruption of the delicate balance of our microbiota can lead to a whole range of health problems, such as inflammatory bowel disease, obesity, colitis, cancer, bacterial vaginosis, strep throat, ear infection, nose congestion, eczema, chronic fatigue syndrome, and more. This phenomenon is known as dysbiosis.
Since 2012 commercial companies have made it possible for anyone to test their microbiome. Among them are the American Gut Project (available only in the U.S. and testing only the gut microbiome), uBiome (available worldwide and testing any site your want) and Atlas Biomed (available in the UK, Netherlands, Belgium and Sweden). Unfortunately at present these companies do not offer much information about the role and significance of the hundreds of bacterial species they identify. I hope with this article to help people understand better what their results mean.
Friend or foe? The role of bacteria in diseases
Properly speaking bacteria do not live in our bodies, but rather on our bodies, on our skin and in mucosal cavities that pass through our body, like the respiratory and digestive tracts where air and food passes. Bacteria should never penetrate the bloodstram or within the organs themselves. Any bacteria is potentially pathogenic if it enters the bloodstream or set up residence within our organs. When this happens, however, our immune system reacts immediately and gets rid of the invaders. The term pathogenic bacteria is used for species that are so aggressive that they to kill commensal bacteria, leading to dysbiosis (e.g. diarrhea), or manage to defeat or thwart or white blood cells, causing inflammation and tissue damage (think pneumonia, sinusitis, otitis, meningitis and the like).
Most, if not all, autoimmune diseases are also caused by bacteria (or more rarely fungi) that manage to infiltrate the body and that the immune system cannot get rid of, either because we are too weak (immunocompromised, overly tired, fighting too many infections at once), or our immune system (HLA system) cannot recognize and appropriately fight one type of pathogen, or because damage to the gut lining (known as Leaky Gut Syndrome) caused by environmental toxins, food intolerances, excess alcohol, antibiotics and the like, allow bacteria to penetrate in the body.
Once they are inside, they will set up colonies in areas difficult to reach by the immune system (i.e. with low blood flow) like the joints, causing autoimmune conditions such as rheumatoid arthritis or ankylosing spondylitis. The latter is caused by the Klebsiella bacteria and typically invades host who possess the HLA-B27 type. Other bacteria might infect the pancreas, causing Type I diabetes, or the muscle fibre, causing fibromyalgia, or directly attacking nerves, which first causes peripheral neuropathies then can lead to Multiple sclerosis (MS). Chronic Fatigue Syndrome is often caused by infections of extremely tiny and primitive bacteria that lack a cell wall known as Mycoplasma, or other unusual bacteria like Borrelia (the cause of Lyme Disease). It has been reported (by Dr Schoemaker) that some types of HLA-DR types cannot easily get rid of Borrelia (DR15, DR16), Dinoflagellates (DR4) and other types of bacteria.
Additionally, bacteria (and fungi) can form a biofilm around them, a sort of slime that protects them from the immune system. If a body part is infected by a bacterial colony protected by a biofilm it will cause a chronic disease that can last for years or even for one's life time. New medicine are being developed to dissolve biofilms, and some natural supplements like serrapeptase could achieve just that.
The interaction between the immune system and our microbiota is what keeps us healthy and makes us sick. That is possible the single most important thing to know about one's health, and yet until recently it was impossible to know what bacteria lived on our bodies. Most people don't know their HLA types (even doctors) even though they are far more useful than blood types or commonly prescribed tests like cholesterol levels. 23andMe uses HLA types to determine risks for dozens of medical conditions, from psoriasis to Type I diabetes.
Gut microbiome
First, the basics. There are five main phyla of bacteria found in the human digestive tract:
Firmicutes : The most common phylum in most Westerners. It includes both beneficial digestive bacteria (Anaerostipes, Blautia, Dorea, Flavonifractor, Lactobacillus, Pseudobutyrivibrio, Robeburia, Ruminococcus, Sarcina, etc.) and pathogenic ones (Listeria, Staphylococcus, Clostridium difficile). Depending on their class, they can be either Gram-positive or Gram-negative.
Bacteroidetes : The second most common phylum among Westerners, but the most common in some non-Western countries. Bacteroidetes are Gram-negative rod-shaped bacteria that can help us digest vegetables (Prevotella), digest fats (Alistipes), digest whole grains and ferment glucose (Bacteroides), or help us regulate our immune system (Barnesiella). Most other Bacteroidetes, however, are pathogenic and shouldn't be found in a healthy microbiome. For example, Porphyromonas have been linked to rheumatoid arthritis, periodontal disease and bacterial vaginosis.
Verrucomicrobia : Tiny wart-shaped bacteria usually found in fresh water and soil. The most common species found in the human gut is Akkermansia, which degrades the excess mucin (mucosal slime) produced by the gut lining. Akkermansia is anti-inflammatory and protective against obesity, colon cancer and autism.
Proteobacteria : These Gram-negative bacteria are essentially bad and include lots of dreaded pathogens like : Bordetella (pertussis), Brucella (brucellosis), Campylobacter (gastroenteritis), Escherichia coli, Helicobacter pylori (gastritis and gastric ulcers), Salmonella, Shigella, Vibrio (e.g. V. cholerae, the causative agent of cholera), and Yersinia (e.g. Y. pestis, the causative agent of the plague). The varieties most commonly found in the gut are Enterobacter (opportunistic infections), Haemophilus (common in the mouth and nose, but can cause respiratory tract infections), Desulfovibrio (linked to IBS and autism), Kluyvera, Sutterella (linked to autism) and Thalassospira.
Actinobacteria : Some of the most common microbes found in our mouths and genitals. They are Gram-positive and typically play a role in decomposing organic matter. The varieties found in the gut are usually beneficial like Collinsella, Cryptobacterium curtum and Bifidobacterium (which is even used as a probiotics and added to Activia yoghurt).
What proportion of good bacteria do you want in your gut ?
Brown et al. (2011) explained how butyrate-producing bacteria protects your gut from inflammation, ulcerative colitis and colorectal cancer. Six main families of firmicutes are known for their ability to convert lactic acid into butyric acid (butyrate). These are Anaerostipes, Flavonifractor, Faecalibacterium, Pseudobutyrivibrio, Roseburia and Subdoligranulum. Butyric acid induces mucin synthesis and tightens the junctions between epithelial cells, thus preventing inflammation and leaky gut syndrome.
Nevertheless, Bacteoridetes like Bacteroides and Alistipes will convert lactic acid into other short-chain fatty acids (SCFAs) like acetic acid, formic acid or propionic acid, which, if present in too large quantities, will damage the lining of the gut, causing inflammation and hyperpermeability of the intestines, leading to autoimmune diseases. So, although Bacteroides and Alistipes are useful and beneficial to digest whole grains and fats, if their proportion exceeds that of the butyrate-producing firmicutes above, it will most probably cause illness. It is therefore important to keep a higher ratio of butyrate-producing bacteria - if possible two or three times more than the Bacteroides and Alistipes. But you also don't want to have too few Bacteroides and Alistipes, as they can also protect you against pathogenic bacteria.
Lactic acid is produced by the lactic bacteria found in yoghurt and probiotics like lactobacillus and bifidobacterium.
Here is the chart summarising the two pathways (from Brown et al. 2011).

Methanobrevibacter smithii is fairly unique but useful bacterium. It is the only member of the Euryarchaeota phylum commonly found in the human gut. It has the capability to reduce sulfate and to recycle excess hydrogen and carbon dioxide into methane, which allows an increase in the extraction of energy from nutrients by shifting bacterial fermentation to more oxidized end products. In other words M. Smithii optimizes energy intake, which causes people to eat less. As a result M. Smithii was found at higher frequencies in lean than in obese people, while anorexics had even higher levels.
I have made a table summarising the main commensal gut bacteria and their function, as well as other useful details like the food they digest.
Role of gut bacteria in diseases
Numerous studies have linked gut dysbiosis to Inflammatory Bowel Disease (IBD). This generally involved a excessive ratio of Bacteroidetes to Firmicutes, but also the presence of specific pathogenic bacteria like Escherichia coli (Darfeuille-Michaud 2004), Mycobacterium avium (Rosenfeld and Bressler 2010), Klebsiella pneumoniae and Proteus mirabilis (Garrett et al. 2010). Some species appear to have a protective role against IBD, like Faecalibacterium prausnitzii and other butyrate producers.
Crohn's Disease patients were found to have more Ruminococcus gnavus and Collinsella aerofaciens but less Ruminococcus torques than healthy subjects or unaffected relatives (Joossens et al. 2013).
Celiac disease patients also showed a greater diversity of Bacteroidetes and Prevotella species than healthy controls (De Palma et al. 2010, Schippa et al. 2010) and had significantly reduced levels of Bifidobacterium and Faecalibacterium.
Gut dysbiosis appear to play an important role in autism. Autistic children were found to have more Bacteroidetes (Finegold et al. 2010) and Clostridium species (Finegold et al. 2002, Parracho et al. 2005)) in their stools, and in the latter case particularly of C. histolyticum. Wang et al. 2013 reported an increased abundance of Sutterella wadsworthensis and well as of Ruminococcus torques and Ruminococcus gnavus in feces of children with autism spectrum disorder who also suffered from constipation and diarrhea. Sutterella is generally absent or represents less than 1% of the gut flora of children who do not have autism, but is almost always present at frequencies of 1 to 7% in autistic individuals. Wang et al. 2011 also found a lower relative abundances of the mucolytic bacterium like Akkermansia muciniphila and of Bifidobacteria in feces of children with autism.
Bifidobacteria, found in many probiotics, appear to play a protective role against several autoimmune diseases. In addition to Celiac disease, patients with rheumatoid arthritis (Vaahtovuo et al. 2008) and diabetes (Wu et al. 2010) were found to have significantly reduced levels of Bifidobacteria.
The microbiota of elderly people was found to be characterized by low species-level diversity. Although great variations were observed between individuals, Claesson et al. 2010 reported that elderly people generally display a decrease in the ratio of Bifidobacteria and Firmicutes but enriched in Bacteroidetes. Biagi et al. 2010 found that centenarians had a spike in undesirable Proteobacteria and Bacilli (e.g. Staphylococci), a likely sign that the immune system cannot get rid of pathogens any more.
Proteobacteria include most of the pathogenic species to human health. A study by UCLA scientists (Clarke et al. 2015) showed that the sudden increase in intestinal permeability (leaky gut) and ratio of gammaproteobacteria (little buggers like Enterobacter, Escherichia coli, Klebsiella, Pasteurella, Pseudomonas or Salmonella) is a sign of rapid health decline and imminent death in fruit flies. The same could apply to humans. Affected fruit flies treated with antibiotics survived three times longer (20 days, which is the equivalent of 30 human years). The study shows that age-onset decline is very tightly linked to changes within the community of gut microbes.
Mouth microbiome
The oral microbiome contains over 800 strains of bacteria. However, Ubiome doesn't test beyond the genus level, so most people will have anywhere between 15 and 100 types of bacteria at the genus level. At the phylum level, Firmicutes and Proteobacteria tend to dominate.
The most common is usually Streptococcus, which has over 50 species and includes the first colonisers of the enamel of the teeth after brushing, strains such as S. oralis, S. mitis, S.gordonii and S. sanguis. Streptococci often make up about half of the oral microbiome. A few species are pathogenic, like Streptococcus pyogenes causing strep throat infections, or of Streptococcus mutans and Streptococcus sobrinus causing dental caries. But many of neutral or even beneficial, like Streptococcus salivarius, which is used as dental probiotics.
Mouth bacteria associated with a good oral hygiene include (most species of):
- Fusobacterium
- Gemella
- Neisseria (although the genus include two famous pathogens: N. meningitidis and N. gonorrhoeae, which do not normally live in the mouth)
These bacteria are part of the normal oral biota and thought to be rather benign or neutral:
- Abiotrophia
- Granulicatella
- Haemophilus (but some species like H. influenza can cause ear, throat and lung infections)
- Leptotrichia buccalis
- Veillonella
It is not unusual to find intestinal bacteria making their way until the oral cavity, although these will be found in low frequencies (<1%) and often barely at trace frequencies (0.01%). Elevated levels of gut bacteria in the mouth may be a sign of Gastroesophageal reflux disease (GERD), linked to excessive gastric acids. Conversely, the presence of more than 0.1% of mouth bacteria in the gut (e.g. Gemella, Veillonella, Haemophilus) could be a sign of insufficient gastric acidity (usually due to ageing), which can lead to nutritional deficiencies through decreased absorption.
Bacteria associated with dental caries (i.e. transforming sugars into lactic acid include mostly:
- Streptococcus mutans
- Streptococcus sobrinus
- Some Lactobacillus species, including the common gut probiotics Lactobacillus acidophilus (which fortunately has low affinity for tooth surfaces)
- Actinomyces viscosus
- Nocardia
- Staphylococcus aureus (perhaps the most troublesome bacterial species for humans)
Note that dental plaque is a kind of bacterial biofilm.
Many more bacteria are known to cause periodontal diseases such as gingivitis or periodontitis.
- Aggregatibacter actinomycetemcomitans
- Campylobacter
- Filifactor
- Fusobacterium nucleatum
- Fusospirochetal gingivitis
- Porphyormonas gingivalis (but not all other porphyromonas species)
- Prevotella
- Tannerella forsythia
- Treponema denticola
Bacteria associated with halitosis (bad breath) include:
- Bacteroides loescheii
- Centipeda periodontii
- Eikenella corrodens
- Enterobacteriaceae
- Eubacterium
- Fusobacterium nucleatum nucleatum
- Fusobacterium nucleatum polymorphum
- Fusobacterium nucleatum vincentii
- Fusobacterium periodonticum
- Porphyromonas endodontalis
- Porphyromonas gingivalis
- Prevotella intermedia
- Prevotella melaninogenica
- Tannerella forsythensis (Bacteroides forsythus)
- Treponema denticola
Skin microbiome
There are about 800 bacterial species found on the skin. They belong essentially to the same phyla as gut bacteria: firmicutes, bacteroidetes, actinobacteria and proteobacteria, although actinobacteria are much more common on the skin.
Acne is caused by Propionibacterium acnes, although usually not all by itself but in interaction with other bacteria like Staphylococcus aureus or Staphylococcus epidermidis.
Eczema is caused by an overgrowth of Staphylococcus aureus, which grows when the skin is slightly alkaline. S. aureus is the principal cause of many skin infections like boils, impetigo, atopic dermatitis, and cellulitis. Left untreated some of these infections (especially pus-filled boils) can invade the blood stream, cause severe fever and even septicaemia and death. The best protection against S. aureus is Staphylococcus epidermidis commensal to human skin and which prefers slightly acidic environments (pH 4-5). So if you suffer from one of the above skin infection, do not use alkaline soaps (including traditional bar soap and Marseilles soap), creams or cosmetics. Some body soaps and most shampoos are slightly acidic, although it is rarely mentioned on the bottle.
Beware that alcohol, which is found in antibacterial lotions used in hospitals but also in most skin care products, damages the natural antibacterial barrier of the epidermis by increasing skin permeability. The reason alcohol is added to skin lotions and creams is that it helps the penetration of ingredients like vitamin A and C. But it also increases the chances of S. aureus infections. That's probably not worth the risk. Alcohol is also slightly alkaline, so its use as an antibacterial will anyway disturb the normal (acidic) skin pH and favour the recolonisation of the skin by S. aureus afterwards.
Few doctors realise that disinfecting skin with alcohol before surgery actually increases the chances of invasive bacterial infections after surgery, once the alcohol has evaporated, but the epidermal junctions have been compromised and alkalised. That is why skin disinfectants should logically be on the acidic side (e.g. lemon juice diluted in water). It's the same problem as with antibiotics that increase gut permeability and lead to bacteria invading the body and causing autoimmune diseases. We are only starting to realise that some medicines, be them alcohol-based disinfectants or antibiotics can do as much harm as good. The problem is that researchers are realising it, but doctors usually fail to catch up with the research (few have the time or incentives to do it). Antibiotic resistance and hospital infections are the result of doctors not caring enough about the microbiome. This unfortunately results in the death of hundreds of thousands of people each year. Hospital infections kill 75,000 people in the US alone every year, and the US population makes up less than 5% of the global population.
Nose & Ear Microbiome
The nasal microbiota is less diverse than in the mouth and gut. It is intermediary between the mouth and skin in terms of species found. Oral bacteria like Bacteroides, Gemella, Veillonella, Lachnospira, Streptococcus and Lactobacillus often end up in small quantity in the nose as do skin bacteria like Corynebacterium, Propionibacterium and Staphylococcus. Among them, Staphylococci are usually the most common, and can indeed represent the vast majority of nasal bacteria.
Staphylococcus aureus, Streptococcus pyogenes and Streptococcus pneumoniae are known to cause respiratory infections.
The nose in fact hosts a lot of not so nice bacteria, especially proteobacteria like Haemophilus, Moraxella, Klebsiella and Pseudomonas, which all include pathogenic species linked to upper respiratory infections. Not all species are pathogenic, however, so their presence in your uBiome results don't mean much without knowing the species involved. Unfortunately uBiome does not test species within a genus at the moment, so it has little diagnostic value for respiratory infections.
One of the bacterium most specific to the nasal cavity is the more benign Dolosigranulum, a Firmicute of the Carnobacteriaceae family, which is not normally found in the skin or mouth.
The ears are connected to the nasopharynx via the Eustachian tube, so that pathological nose bacteria will also generally infect the ear and cause otitis media. The most common bacteria causing middle ear infections are Streptococcus pneumoniae, Haemophilus influenzae, Moraxella catarrhalis and Staphylococcus aureus.
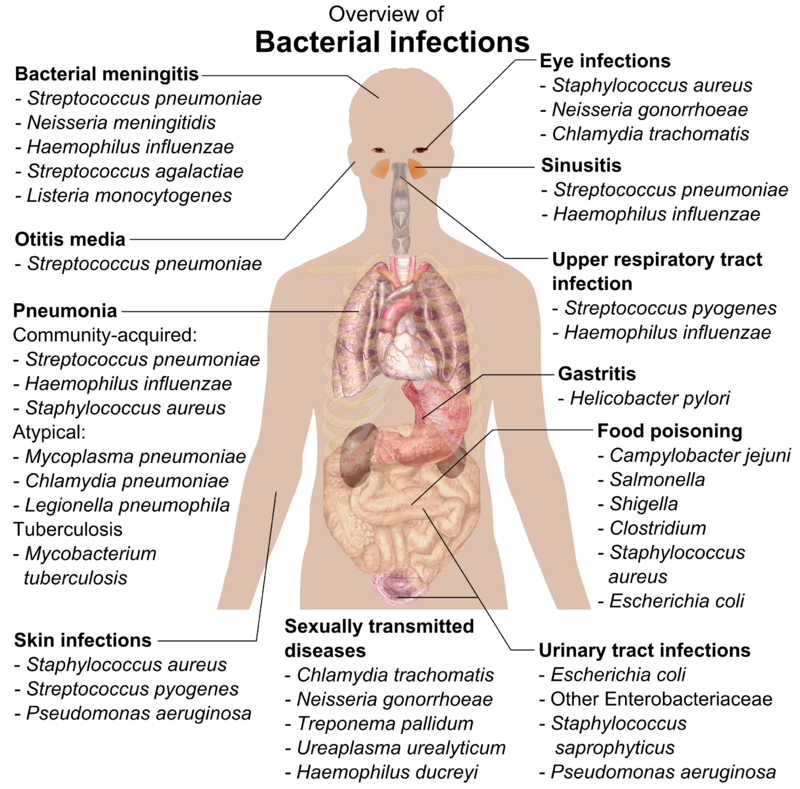
Here is a chart of common pathogenic bacteria from Wikipedia.
The bottom line: are microbiome tests useful yet?
Microbiome tests are still in their infancy. The human microbiota is made of 100 trillion bacterial cells composed of thousands of species of bacteria, and many more subspecies, but also numerous species of fungi, yeasts and viruses, some harmful, others harmless or even beneficial. Quite a few bacterial species cannot easily be categorised as good or bad for health, as some subspecies are benign or favourable, while others can be pathogenic and, in some cases, even deadly. That is the case of well known species, such as Staphylococcus and Streptococcus. Unfortunately, tests like uBiome do not distinguish between subspecies yet, which make the results pretty useful for such ambivalent species. Will the Streptococci in one's mouth be the strep throat causing S. pyogenes, the S. sobrinus causing dental caries, or the beneficial S. salivarius that protects against both caries and and strep throat infections? The results won't tell you.
Additionally, the uBiome Explorer test only analyse bacterial species, not fungi (like Candida) or viruses that also play an primordial role in one's microbiota and overall health. Another drawback is that too little is known about most bacterial species, their role in various diseases, their interactions with one another, the role of the immune system (and notably HLA types) in controlling or eliminating specific bacterial species, and the real long-term effect of probiotics on the microbiome. Besides, exisitng probiotics only belong to a handful of bacterial species such as Lactobacillus and Bifidobacterium, and therefore cannot replenish other beneficial species that may be deficient. Our understanding of the influence of nutrition of the microbiome is patchy at best and cannot be used to properly rebalance one's microbiome.
Finally, current antibiotics are not able to target specific bacterial species or subspecies. Although a new generation of species-specific treatments (including nanobots) is under development, it may be years or decades before tailored treatments become available to kill a precise pathogenic species. In conclusion, although the concept of microbiome testing is attractive, the current knowledge and depth of testing are not sufficient to provide any substantial therapeutic benefits.

